CometChip: Microarray for DNA Damage
 For decades, researchers have measured DNA damage using the single cell gel electrophoresis assay, or comet assay. Originally developed by Singh et al in 1988, the basis for the assay is that damaged DNA migrates more readily than undamaged DNA when electrophoresed. Although effectively used in thousands of publications, the assay nevertheless suffers from sample-to-sample variation and it is highly laborious.
For decades, researchers have measured DNA damage using the single cell gel electrophoresis assay, or comet assay. Originally developed by Singh et al in 1988, the basis for the assay is that damaged DNA migrates more readily than undamaged DNA when electrophoresed. Although effectively used in thousands of publications, the assay nevertheless suffers from sample-to-sample variation and it is highly laborious.
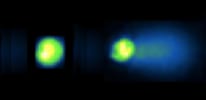

Increasing amounts of DNA damage are visible in the comets below. The nucleoid from a normal cell is on the left, whereas the nucleoid from a cell exposed to DNA damage is on the right.
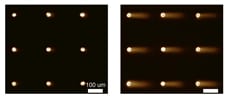
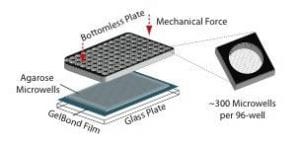
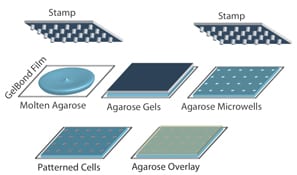
To increase reproducibility and throughput, we collaborated with the Bhatia laboratory (MIT HST EECS). Using microfabrication, we developed a method for making an array of microwells in agarose. A single cell (or groups of cells) can be loaded into each well.
Traditionally, 100 comets are analyzed for each sample, and it had been necessary to image each comet individually, which was highly laborious. By creating a cell microarray, comets share the same focal plane, making it possible to capture 100 comets in just one, saving hours in labor. In addition, in-house software enables automated comet analysis, which reduces bias and increases throughput.
The platform is useful for studies of single cells or clusters of cells, and it has been shown to be effective for analysis of DNA damage in a broad range of mammalian cell types, from human primary cells to human cancer cells.
|
|